What Are Nanodiscs?
The term nanodiscs describes a small (7-50 nm in diameter) disc-shaped structure that finds use in proteomics and biomedicine. It consists of two main components:
- Phospholipids, either of the artificial origin or from the cell membrane
- A stabilizing belt that holds the phospholipids together. This is either an MSP or a synthetic polymer.
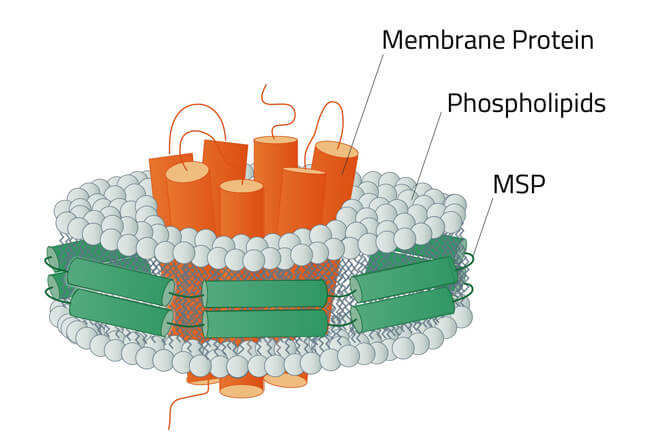
Green: The stabilizing belt - A MSP in this case
Grey: Phospholipids
Orange: Stabilized membrane protein
The Purpose of Nanodiscs
Their aim is to mimic the native phospholipid bilayer of cells for target molecules (often membrane proteins). Membrane proteins are the key to communication between cells. They mediate fundamental biological processes such as signal transduction, transport processes across membranes, sensing of chemical signals, and coordination of cell-cell interactions.
Numerous diseases in humans are linked to membrane proteins, making them important targets for drug development. So it seems surprising, that membrane proteins are encoded by up to ∼23% of genes but represent <1 % of known protein structures (1).
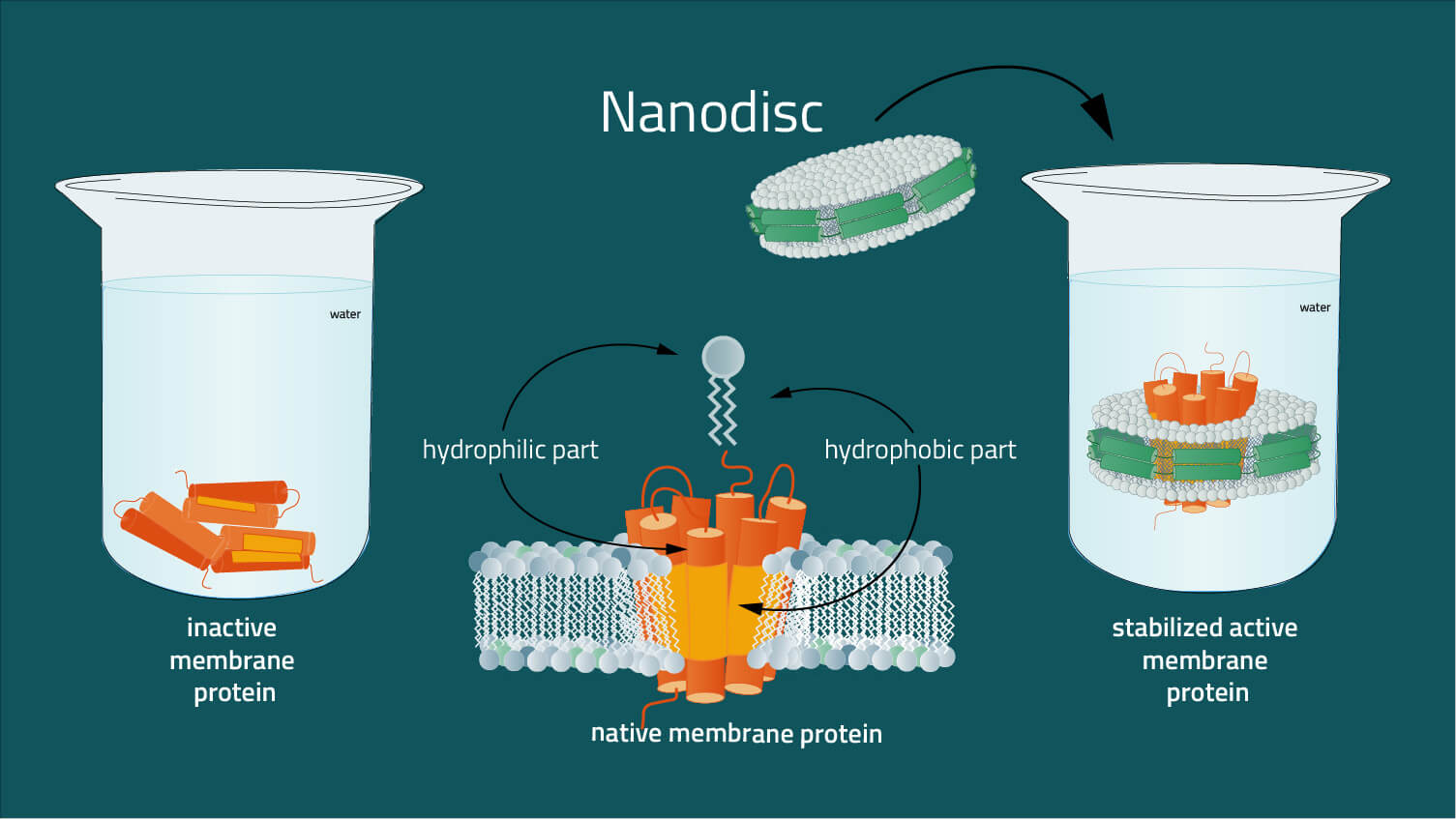
The problem is, that membrane proteins, unlike soluble proteins, are difficult to analyze in their native environment, due to their insertion in the lipid membrane. Surfaces of membrane proteins in the hydrophobic core of the lipid bilayer are also hydrophobic, whereas surface areas in contact with the aqueous membrane environment are as hydrophilic as the surfaces of ordinary soluble proteins (2). The presence of extensive hydrophobic and hydrophilic surfaces on the same molecule is characteristic of membrane proteins. As a result, membrane proteins are not soluble in standard aqueous buffers without a solubilizing agent. For example, nanodiscs are required to solubilize them to mimic the amphipathic environment of a lipid bilayer whilst maintaining the structure of the membrane protein in a physiologically relevant state.
Watch our "The purpose of nanodiscs" video guide, for an animated summary of this chapter.
Two Types of Nanodisc
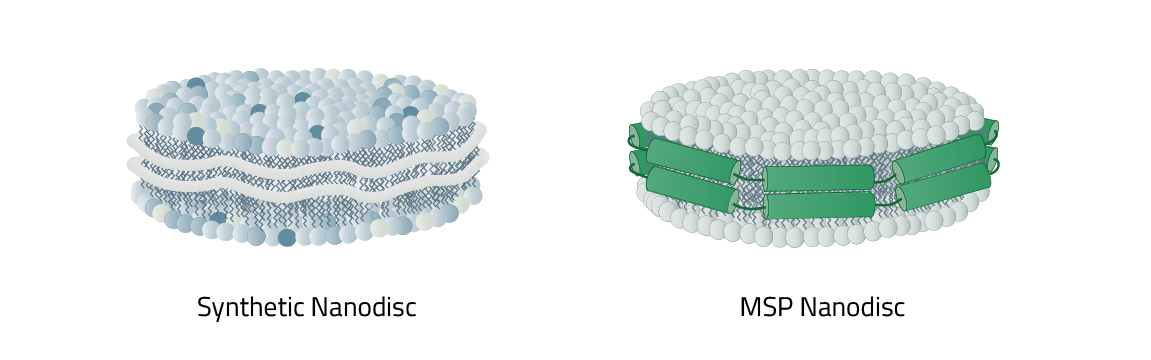
As mentioned before nanodiscs can be differentiated between their phospholipid composition and most importantly, their type of stabilizer. This stabilizer is the reason why nanodiscs in total are split into two main categories: MSP nanodiscs and Copolymer Nanodisc.
The respective names originate from the type of stabilizer that is used to keep the nanodiscs together and form them in the first place. It also decides what the lipid composition of the nanodisc is made up of. MSP nanodiscs always contain an artificial lipid composition. Meaning you have full control of it. In contrast: Copolymer Nanodisc use the native cell phospholipids to create the nanodisc. A direct comparison of the individual advantages of both nanodiscs can be found here.
Table 1: Small comparison between MSP and Copolymer Nanodisc
| Copolymer Nanodisc | Membrane scaffold protein (MSP) Nanodisc | |
|---|---|---|
| Belt properties / Stabilizer | synthetic Copolymer (e.g. DIBMA, SMA) | MSP |
| Lipids | Native cell membrane lipids | Artificial phospholipid environments (e.g. phospholipids) |
| Examples | Diisobutylene-maleic acid (DIBMA), styrene-maleic acid (SMA), amphipols | Pre-Assembled nanodiscs |
But first, let us introduce both kinds of nanodiscs on their own, starting with the MSP nanodiscs.
MSP Nanodiscs
MSP nanodiscs are held together by membrane scaffold proteins (MSPs). MSPs can be truncated forms of apolipoprotein (apo) A-I which wrap around a patch of a lipid bilayer to form a disc-like particle or nanodisc (5). MSPs provide a hydrophobic surface facing the hydrophobic tail of the lipids, and a hydrophilic surface on the outside. This setup makes nanodiscs highly soluble in aqueous solutions. Once assembled into nanodiscs, membrane proteins can be kept in solution in the absence of detergents (5).
Size: The size of an MSP nanodisc can range between 7 - 17 nm. It is determined by the used membrane scaffolding protein. Table 2 depicts the membrane scaffold proteins that Cube Biotech offers and which nanodisc sizes it leads to. MSP nanodiscs of the same MSP are uniform in size and only differ +/- 1 nm in diameter. This suits them perfectly for Cryo-EM studies.
Other advantages of MSP nanodiscs
MSP nanodiscs have a number of advantages compared to other systems for membrane protein solubilization and reconstitution, in particular for ligand binding studies, analysis of conformational dynamics, and protein interaction studies (6). Nanodiscs can be used to reconstitute membrane proteins such as GPCRs or transporters in an artificial environment resembling the native membrane.
Size: The size of an MSP nanodisc can range between 7 - 17 nm. It is determined by the used membrane scaffolding protein. Table 2 depicts the membrane scaffold proteins that Cube Biotech offers and which nanodisc sizes it leads to. MSP nanodiscs of the same MSP are uniform in size and only differ +/- 1 nm in diameter. This suits them perfectly for Cryo-EM studies.
Other advantages of MSP nanodiscs
MSP nanodiscs have a number of advantages compared to other systems for membrane protein solubilization and reconstitution, in particular for ligand binding studies, analysis of conformational dynamics, and protein interaction studies (6). Nanodiscs can be used to reconstitute membrane proteins such as GPCRs or transporters in an artificial environment resembling the native membrane.
These nanodisc-stabilized proteins can be directly purified by standard chromatographic procedures. The resulting purified membrane protein-nanodisc complex can be used in applications that require access to both the physiologically intracellular and extracellular surfaces of the protein and thus allows unrestricted access to antagonists, agonists, G proteins, and other interaction partners (7).
How to generate MSP nanodisc + protein - complexes
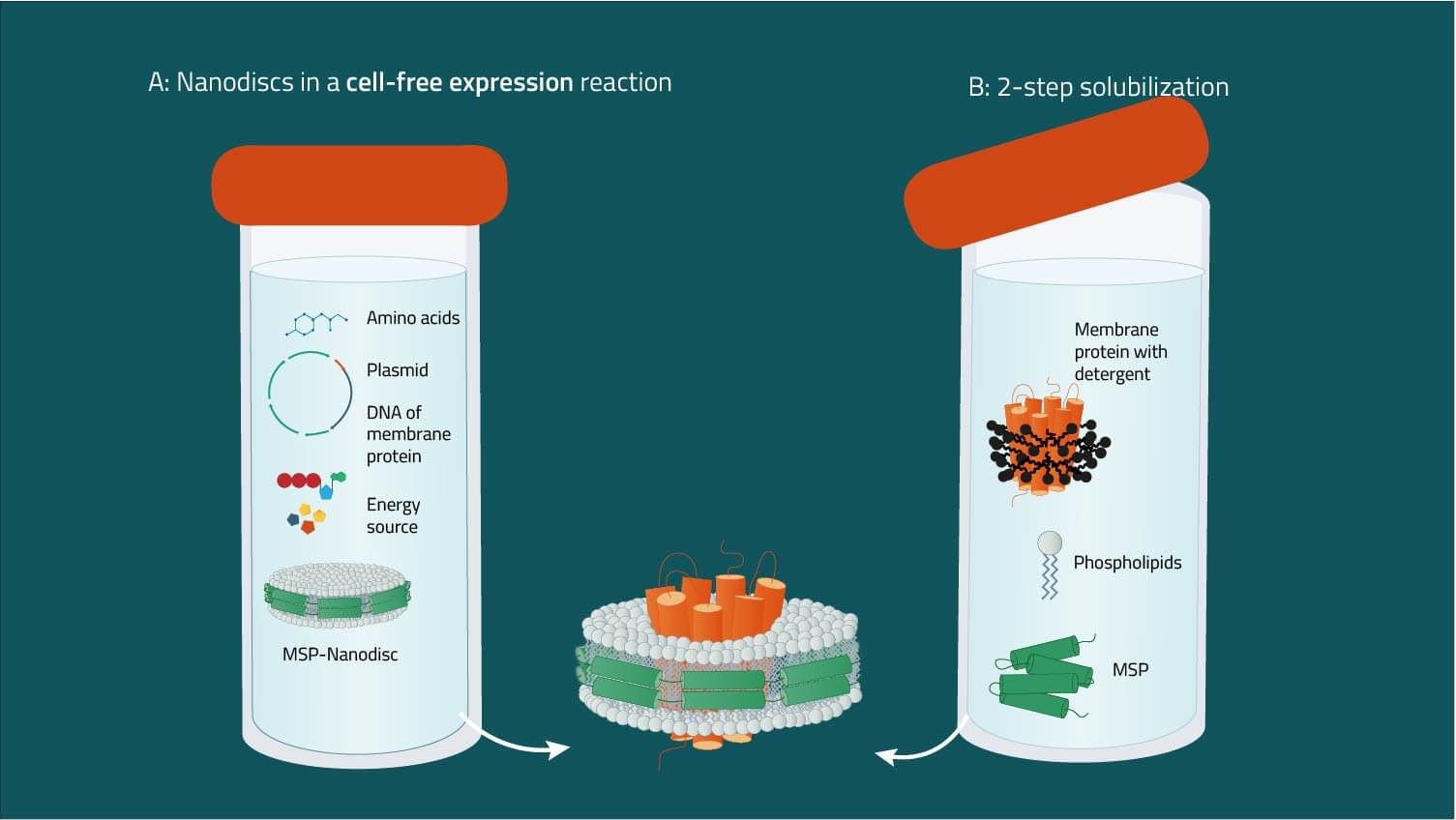
A: Combining nanodiscs and cell-free expression systems
Starting from an expression plasmid, membrane proteins can be produced in cell-free systems. Pre-assembled nanodiscs are supplied in the mixture that integrate the nascent membrane protein (8). The addition of detergents is not required, which minimizes possible artifacts. Optionally, modifications such as biotinylation or isotope labeling can be included.
B: Two-step reconstitution of detergent-solubilized proteins
Starting from a purified membrane protein in suitable detergent, membrane scaffold proteins and phospholipids are added. Nanodiscs containing the membrane protein form spontaneously, and can be purified by affinity or size exclusion chromatography (6, 7).
Starting from a purified membrane protein in suitable detergent, membrane scaffold proteins and phospholipids are added. Nanodiscs containing the membrane protein form spontaneously, and can be purified by affinity or size exclusion chromatography (6, 7).
C: Direct solubilization from membranes
Starting from membranes expressing the protein of interest, detergent and membrane scaffold protein are added. Membrane phospholipids, membrane protein, and MSP assemble to form the nanodisc complex (5). Here, a mixture of nanodisc complexes representing the membrane protein population is obtained, which may be used for proteomics studies. If required, individual membrane protein-nanodisc complexes can be purified by affinity chromatography. Compared to method B, exposure time to detergents is significantly shorter (hours vs. days).
Starting from membranes expressing the protein of interest, detergent and membrane scaffold protein are added. Membrane phospholipids, membrane protein, and MSP assemble to form the nanodisc complex (5). Here, a mixture of nanodisc complexes representing the membrane protein population is obtained, which may be used for proteomics studies. If required, individual membrane protein-nanodisc complexes can be purified by affinity chromatography. Compared to method B, exposure time to detergents is significantly shorter (hours vs. days).
Choice of Phospholipids - the Key to proper Protein Activity
As already mentioned the phospholipids composition of an MSP nanodisc is artificial. Meaning the used phospholipids that should make up the artificial membrane environment for the membrane protein of interest must be decided before. But there are numerous phospholipids to choose from out there, so which to choose? Refer to this list of our most commonly used phospholipids for MSP nanodiscs, when faced with this question.
Dimyristoyl-glycero-phosphocholine (DMPC)
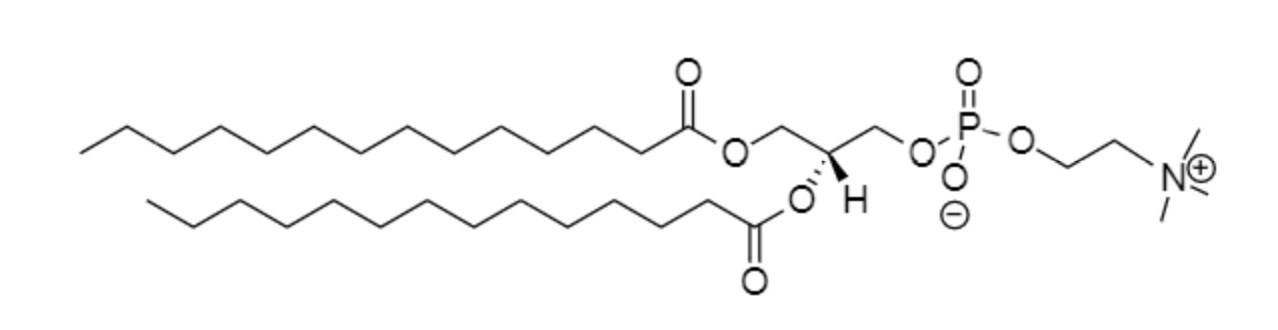
Palmitoyl-oleoyl-phosphatidylcholine (POPC)
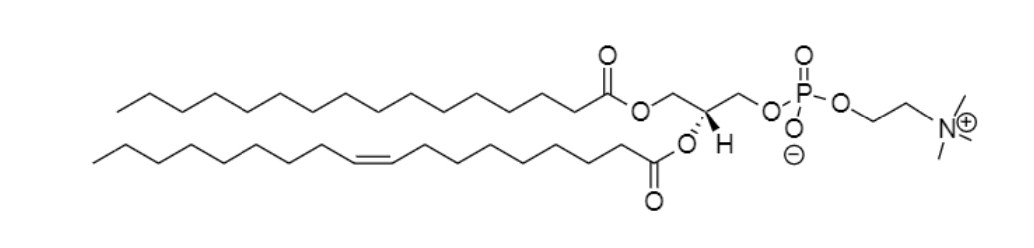
Phosphatidylglycerol (DMPG)
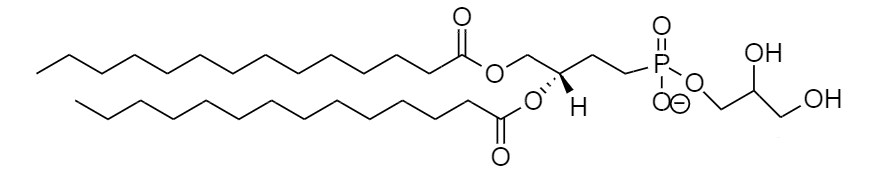
This selection, but also many other phospholipids have been successfully used alone or in combination (8,25). The choice of lipids has been shown to be crucial for protein activity (8), for example in cases where lipids promote protein oligomerization (25). Cell-free expression using assembled nanodiscs is a fast and easy way to screen a variety of lipids and lipid mixtures for their effect on the protein. When proteins are solubilized directly from the membrane fraction, endogenous phospholipids are carried along and incorporated into the nanodisc complex, which may enhance protein activity.
Examples of MSP nanodisc applications in science
MSP Nanodiscs were first described by Sligar and coworkers (3,4). They provide the perfect environment to stabilize membrane proteins to study the binding of ligands, agonists, or antagonists by methods such as NMR and SPR (9,10). Nanodiscs were shown to increase the resolution of membrane-spanning protein regions in Cryo-EM (22,26). Membrane scaffold proteins can be tagged with histidines to facilitate purification, detection, and immobilization of the protein-nanodisc complex. Other nanodisc applications include resonance Raman (11), MALDI (13), non-covalent mass spectrometry (25), protein activation studies (14), time-resolved fluorescence spectroscopy (15), and protein crystallization (24). Antigens reconstituted into nanodiscs have been used to raise immunogenic responses in mice, showing their potential to be used as vaccines (16). In addition, the entire membrane proteome of E.coli was reconstituted into nanodiscs, thereby creating a solubilized membrane protein library (15). Proteins reconstituted in nanodiscs can be transferred to bicelles to improve NMR resolution (23). Even soluble, lipid-interacting proteins were analyzed with the help of nanodiscs (20). Table 3 lists examples for nanodisc applications.
Copolymer Nanodisc
Copolymer Nanodisc are the second big option in the field of nanodisc. They differ in certain key aspects from their MSP counterparts, but also share certain similarities.
Creation of Copolymer Nanodisc
In contrast to the three creation ways of MSP nanodiscs (figure 4), Copolymer Nanodisc can only be created directly from intact cells. The used synthetic Copolymer has a dual function during this process. First, it dissolves the cell membrane, similar to a detergent. Then it forms a nanodisc structure around membrane proteins using the native cell phospholipids. A good analogy to this process is a cookie cutter that stamps the cookies out of the dough.
Creation of Copolymer Nanodisc
In contrast to the three creation ways of MSP nanodiscs (figure 4), Copolymer Nanodisc can only be created directly from intact cells. The used synthetic Copolymer has a dual function during this process. First, it dissolves the cell membrane, similar to a detergent. Then it forms a nanodisc structure around membrane proteins using the native cell phospholipids. A good analogy to this process is a cookie cutter that stamps the cookies out of the dough.
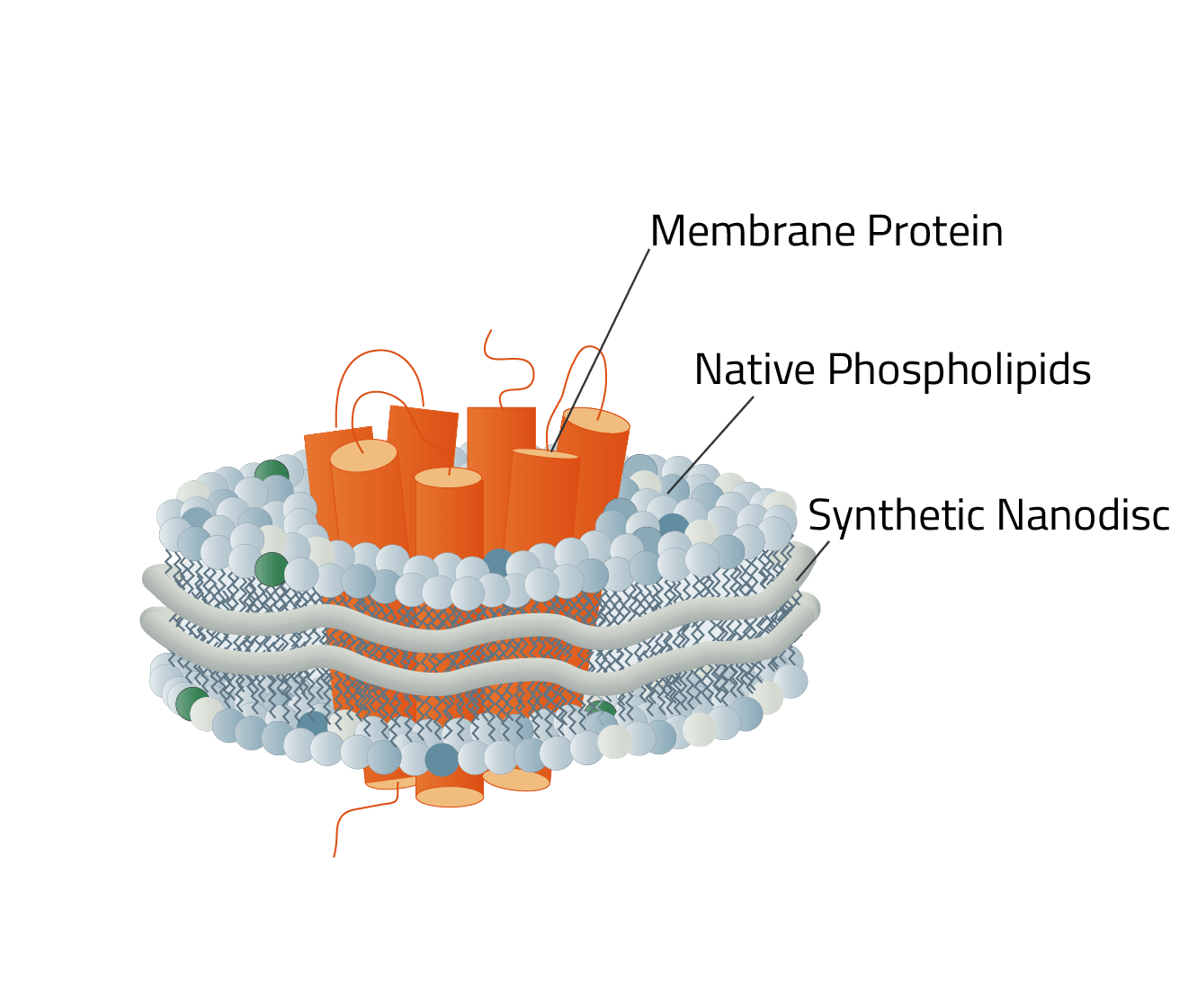
Size
Copolymer Nanodisc are variable in their size. The main factor that decides their diameter is the size of the membrane protein complex that they surround and stabilize. Therefore a definitive size cannot be given for a Copolymer Nanodisc. But they all range in the size range that can also be found in MSP nanodiscs (table 2). This applies to all established Copolymer so far. If uniformous nanodisc size is desired for a Copolymer Nanodisc complex, a size-exclusion chromatography (SEC) has to be performed after the stabilized membrane protein of interest has been purified by e. g. affinity chromatography with the Rho1D4-tag.
Copolymer Nanodisc are variable in their size. The main factor that decides their diameter is the size of the membrane protein complex that they surround and stabilize. Therefore a definitive size cannot be given for a Copolymer Nanodisc. But they all range in the size range that can also be found in MSP nanodiscs (table 2). This applies to all established Copolymer so far. If uniformous nanodisc size is desired for a Copolymer Nanodisc complex, a size-exclusion chromatography (SEC) has to be performed after the stabilized membrane protein of interest has been purified by e. g. affinity chromatography with the Rho1D4-tag.
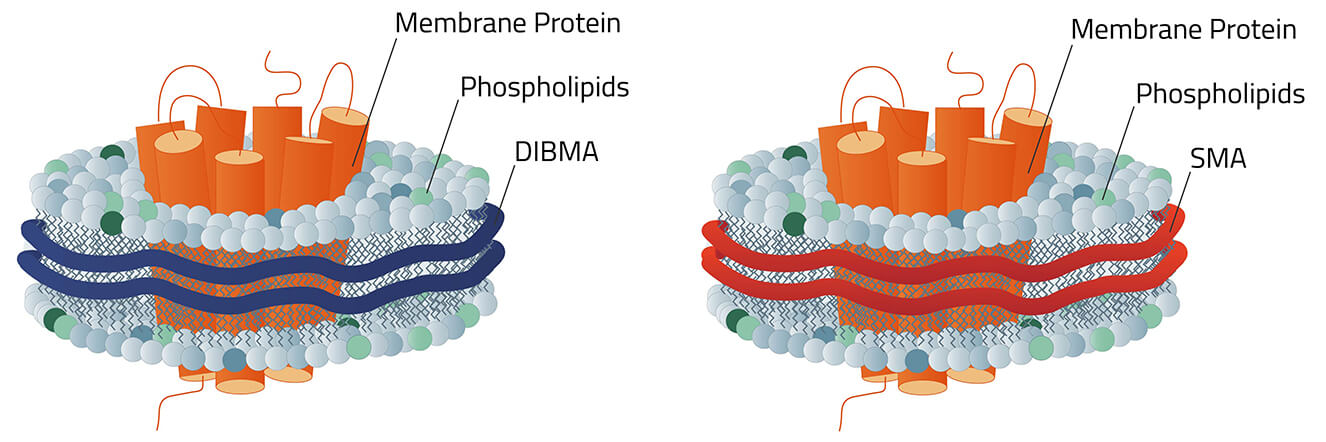
Which Copolymer?
The selection of different Copolymer for Copolymer Nanodisc is constantly growing. Each with its own benefits and downsides. DIBMA for example has protein-like absorption at a wavelength of 280 nm. Meanwhile, AASTY has quite fixed nanodiscs diameters compared to other Copolymer.
However, this is only a small extract from a long list of pros and cons of the different Copolymer. We dedicated this topic its own webpage. Have a look at it!
The selection of different Copolymer for Copolymer Nanodisc is constantly growing. Each with its own benefits and downsides. DIBMA for example has protein-like absorption at a wavelength of 280 nm. Meanwhile, AASTY has quite fixed nanodiscs diameters compared to other Copolymer.
However, this is only a small extract from a long list of pros and cons of the different Copolymer. We dedicated this topic its own webpage. Have a look at it!
More in-depth information regarding the different Copolymer for Copolymer Nanodisc can be found here:
MSP or Copolymer Nanodisc?
So after all of this, the question remains what type of nanodisc best suits your project? Both MSP and Copolymer Nanodisc are meant for the solubilization & stabilization of membrane proteins by mimicking a cell-membrane environment. However, as mentioned before, there are some key differences between the two. Table 4 lists all differences and their respective advantages & disadvantages.
Table 4: Direct comparison between MSP and Copolymer Nanodisc.
| MSP nanodiscs | Copolymer Nanodisc |
|---|---|
| Size Depending on the used MSP. Uniformous (+/-1 nm) for each MSP) Advantage: Uniformous sizes make MSP nanodiscs perfect tools for applications like Cryo-EM. |
Size Variable, due to different lengths of the Copolymer chains. Advantage: The variability of the diameter skips screening steps that are necessary when working with MSP nanodiscs. |
| Lipid composition Artificial. Provided by the scientist. Advantage: The scientist has complete control over the phospholipid composition. |
Lipid composition Made up of native cell membrane lipids. Advantage: The membrane protein is stabilized in a part of its native environment. |
| UV absorption Overlaps with membrane protein due to the presence of the MSP. Note: Due to the MSP, the nanodisc itself has a UV signal at a wavelength of 280 nm and interferes with protein quantification attempts via absorbance. |
UV absorption SMA behaves like MSP nanodiscs, but DIBMA-based nanodiscs do not absorb at wavelengths of 280 nm. Advantage: With DIBMA-based nanodiscs, the protein quantity can easily be determined by measuring the absorbance of the solution at a wavelength of 280 nm. |
| Creation Able to be created in 3 different ways (see figure 4). Advantage: The different situations in which MSP nanodiscs can stabilize membrane proteins make them the go-to option often. |
Creation Only directly from the cell. Note: Since the synthetic Copolymer use native cell membrane material to create the nanodiscs, only membrane proteins from living cell material can be stabilized. |
| Involvement of detergents Involved in the beginning before the MSP form the nanodisc around the protein of interest. Note: A detergent has to be chosen that does not impact the folded protein's structure. This can result in some extra work. |
Involvement of detergents No detergent necessary. Advantage: The Copolymer both act as solubilizers and stabilizers simultaneously. Therefore no additional detergents are needed. |
Literature References
- Douglas, Shawn M., James J. Chou, and William M. Shih. "DNA-nanotube-induced alignment of membrane proteins for NMR structure determination." Proceedings of the National Academy of Sciences 104.16 (2007): 6644-6648.
- Yeates, T. O., et al. "Structure of the reaction center from Rhodobacter sphaeroides R-26: membrane-protein interactions." Proceedings of the National Academy of Sciences 84.18 (1987): 6438-6442.
- Bayburt, T.H. et al. Reconstitution and imaging of a membrane protein in a nanometer-size phospholipid bilayer. J. Struct. Biol. (1998), 123(1):37-44
- Civjan, N.R. et al. Direct solubilization of heterologously expressed membrane proteins by incorporation into nanoscale lipid bilayers. BioTechniques (2003) 35:556-563
- Hagn, F. et al. Optimized phospholipid bilayer nanodiscs facilitate high-resolution structure determination of membrane proteins. J.Am.Chem. Soc. (2013), 135:1919-1925
- Serebryany et al. Artificial membrane-like environments for in vitro studies of purified G-protein coupled receptors. Biochim. Biophys. Acta (2012), 181:225-233
- Leitz, J. et al. Functional reconstitution of beta2-adrenergic receptors utilizing self-assembling nanodisc technology. BioTechniques (2006), 40:601-612
- Proverbio D., et al. Functional properties of cell-free expressed human endothelin A and endothelin B receptors in artifical membrane environments. Biochim.Biophys. Acta (2013), 1828(9):2182-92
- Glueck, J.M. et al. Integral membrane proteins in nanodiscs can be studied by solution NMR spectroscopy. J.Am.Chem.Soc. (2009), 131(34):12060-1
- Glueck, J.M. et al. Nanodiscs allow the use of integral membrane proteins as analytes in surface plasmon resonance studies. Anal. Biochem. (2011), 408(1):46-52
- Mak, P.J. et al. Defining CYP3A4 structural responses to substrate binding. Raman spectroscopic studies of a nanodisc-incorporated mammalian cytochrome P450. J.Am.Chem.Soc. (2011) 133(5):1357-66
- Frauenfeld, J. et al. Cryo-EM structure of the ribosome-SecYE complex in the membrane environment. Nature Struct. Mol. Biol. (2011), 5:614-21
- Marty M.T., et al. Ultra-thin layer MALDI mass spectrometry of membrane proteins in nanodiscs. Anal. Bioanal. Chem. (2012) 402(2):721-9
- Wang, Z. et al. Tyrosine phosphorylation of Mig6 reduces its inhibition of the epidermal growth factor receptor. ACS Chem. Biol. (2013) 8(11):2372-6.
- Pandit A., et al. Assembly of the major light-harvesting complex II in lipid nanodiscs. Biophys. J. (2011) 101:2507-2515
- Bhattacharya, P. et al. Nanodisc-incorporated hemagglutinin provides protective immunity against influenza virus infection. J. Virology (2010) 361-371
- Marty M.T. et al., Nanodisc-solubilized membrane protein library reflects the membrane proteome. Anal. Bioanal. Chem. (2013) 405(12):4009-16
- Moers et al., Modified lipid and protein dynamics in nanodiscs. Biochim. Biophys. Acta (2013), 1828(4):1222-9.
- Nasr. et al., Radioligand binding to nanodisc-reconstituted membrane transporters assessed by the scintillation proximity assay. Biochemistry (2014), 14;53(1):4-6.
- Kobashigawa. et al., Phosphoinositide-incorporated lipid-protein nanodiscs: A tool for studying protein-lipid interactions. Anal. Biochem. 410 (2011), 77-83
- Grinkova, Y.V., et. al., Engineering extended membrane scaffold proteins for self-assembly of soluble nanoscale lipid bilayers. Protein Engineering, Design & Selection (2010) 23(11):843-848
- Gatsogiannis, C, et. al., Membrane insertion of a Tc toxin in near-atomic detail. Nat. Struct. Mol Biol. (2016) Oct;23(10):884-890.
- Laguerre, A. et al. From nanodiscs to isotropic bicelles: A procedure for solution nuclear magnetic resonance studies of detergent-sensitive integral membrane proteins. Structure (2016) 24, 1-12.
- Nikolaev, M. et al. Integral membrane proteins can be crystallized directly from nanodiscs. Cryst. Growth Des. (2017) 17(3), 945–948
- Henrich, E., et al. Analyzing native membrane protein assembly in nanodiscs by combined non-covalent mass spectrometry and synthetic biology. eLife (2017) 6:e20954.
- Gao, Y. et al. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature (2016) 534(7607):347-351. doi:10.1038/nature17964
MSP Nanodiscs are protected by US Patents 7,691,414; 7,662,410; 7,622,437; 7,592,008; 7,575,763; 7,083,958; 7,048,949


