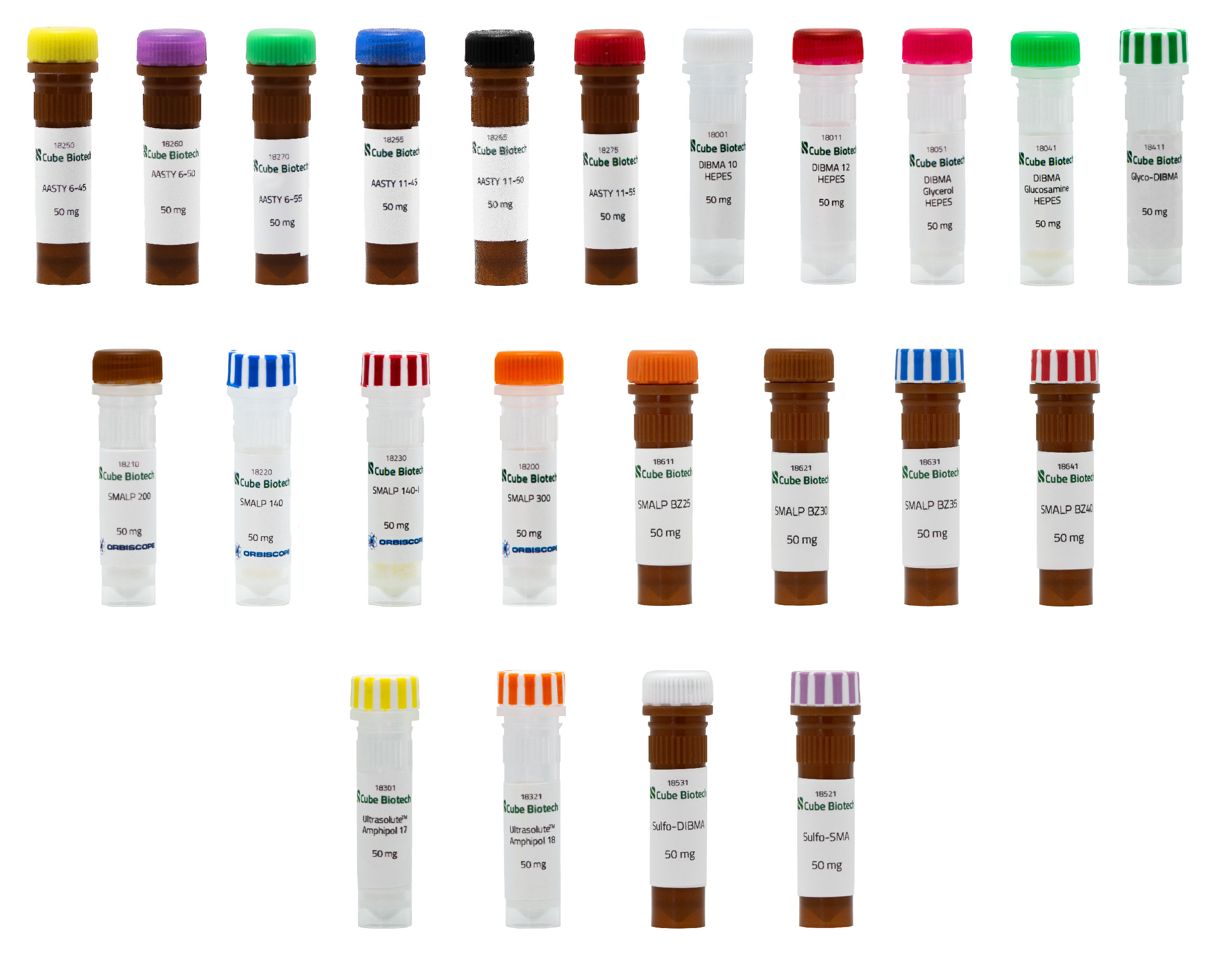
DIBMA
DIBMAs or Diisobutylene-maleic acids are alternatives to detergents which are normally used in membrane protein solubilization even though it is well known that detergents have specific weaknesses. Short-chain nonionic detergents for example can affect the functional properties of a membrane protein. It seems clear that removing the native lipid bilayer from the membrane protein can interfere with the function of the protein. One way to mimic the native lipid membrane are MSP-Nanodiscs or synthetic nanodiscs, like SMAs and the aforementioned DIBMAs. With the latter, you can directly extract membrane proteins from cells without an intermediate step of detergent solubilization. Synthetic polymers have to carry styrene or maleic acid group themselves to solubilize proteins.